Using Moran’s I for assessing residual spatial autocorrelation in machine learning models
2026-05-06, EGU 2026, Vienna
Same RMSE, different prediction maps
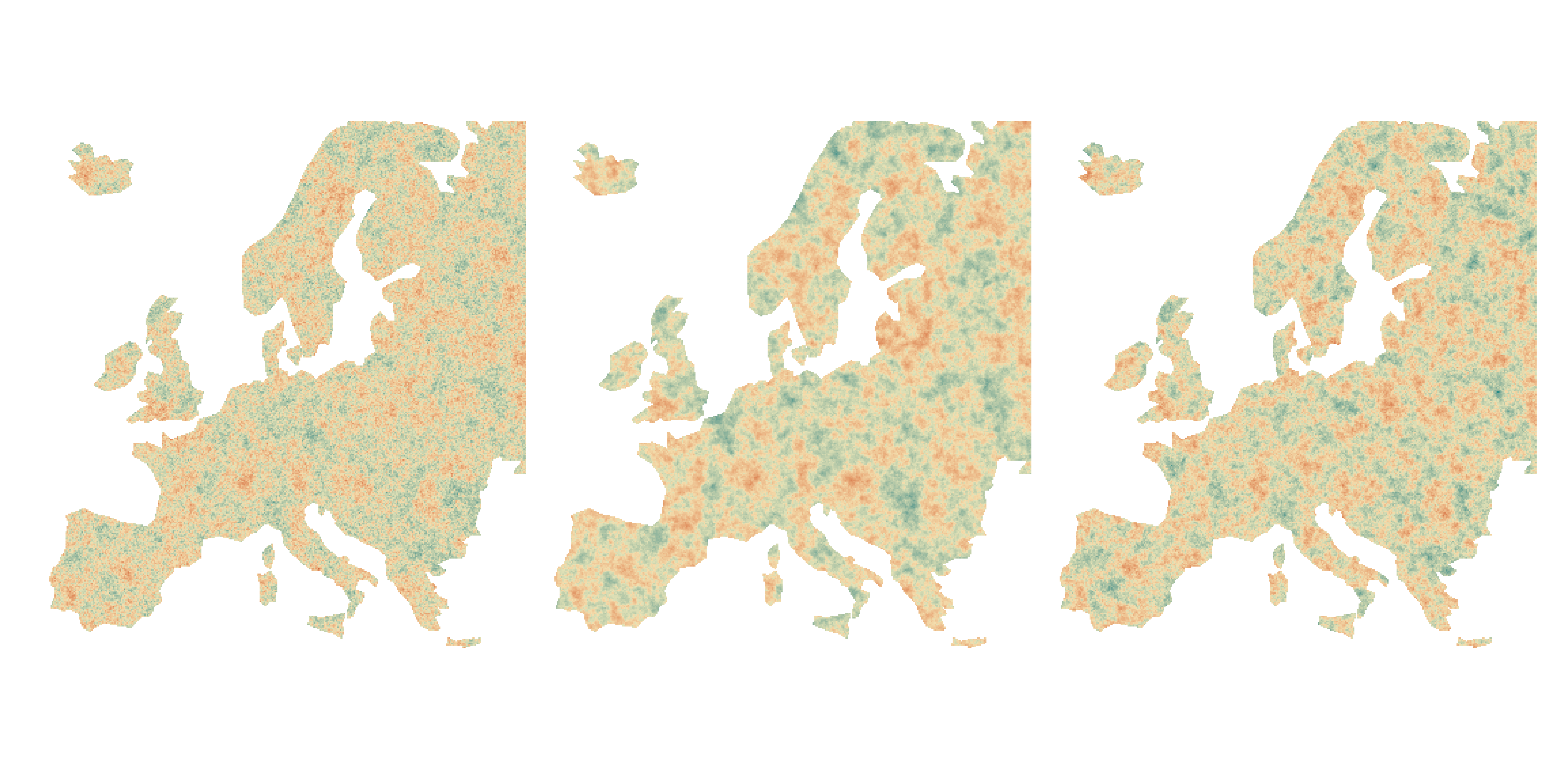
Residuals can still be spatially structured
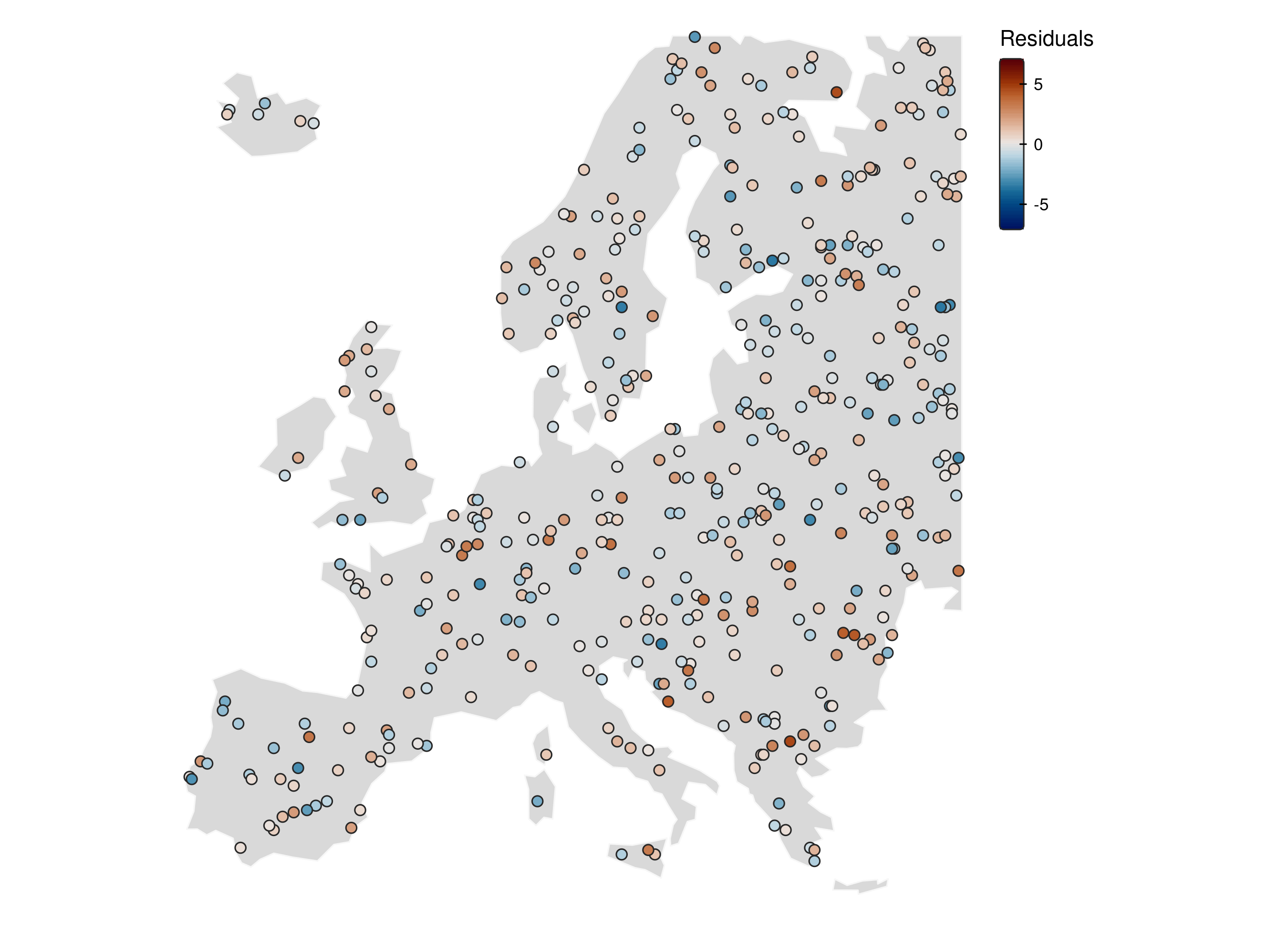
How should we diagnose spatial autocorrelation in model residuals?
Methods
Moran’s I
Moran’s I measures similarity among neighboring residuals
\[ I = \frac{n}{\sum_{i}^{n} \sum_{j}^{n} w_{ij}} \times \frac{\sum_{i}^{n} \sum_{j}^{n} w_{ij} (x_i - \bar{x}) (x_j - \bar{x})}{\sum_{i}^{n} (x_i - \bar{x})^2} \]
- \(n\): number of observations
- \(x_i\), \(x_j\): values of the observations at locations \(i\) and \(j\)
- \(\bar{x}\): mean value of the observations
- \(w_{ij}\): spatial weight between the observations at locations \(i\) and \(j\)
Spatial weight defines which observations are considered neighbors.
kNN-based weights
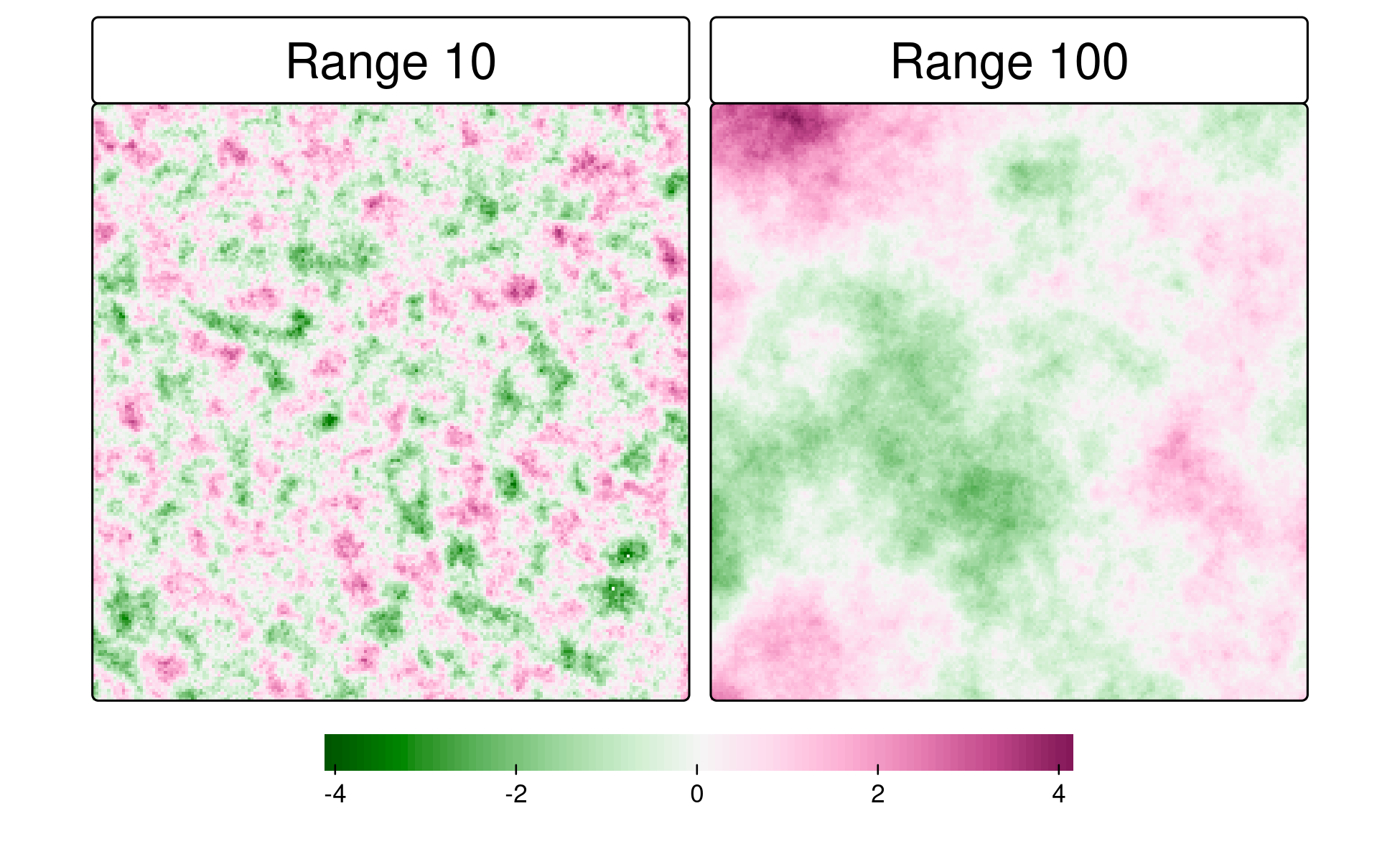
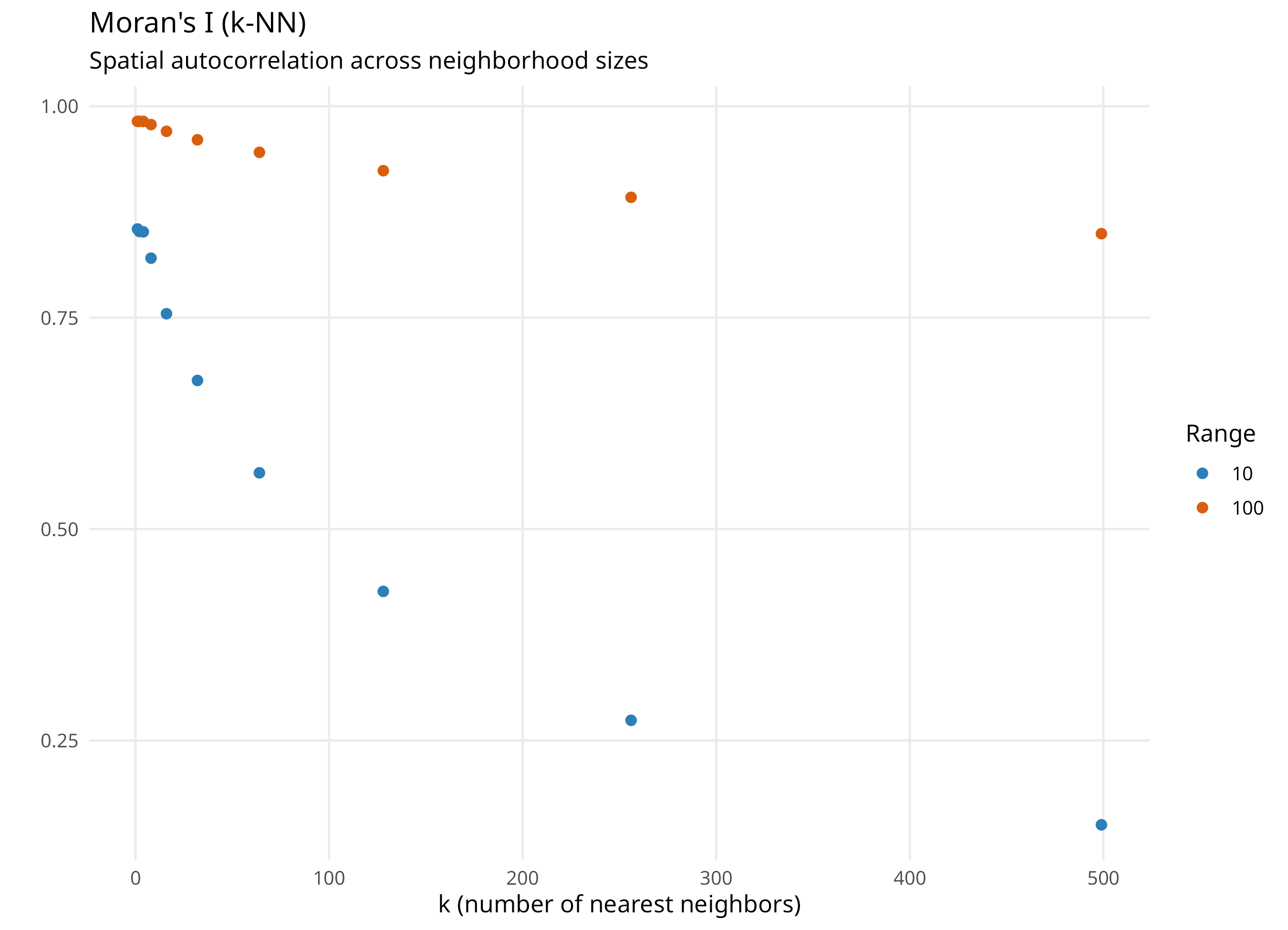
kNN: fixed number of neighbors, variable geographic extent
Larger k captures broader spatial structure
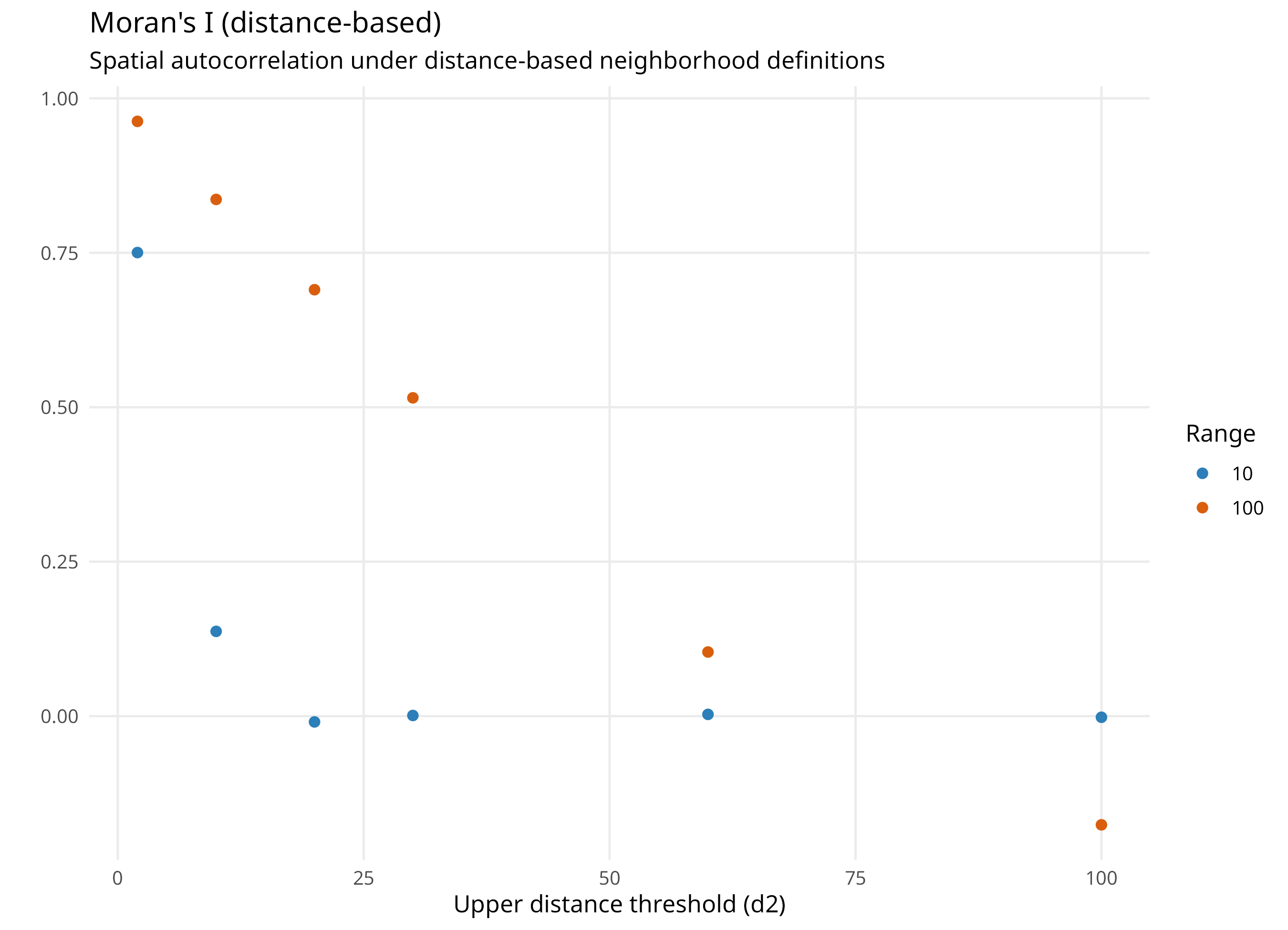
Distance-based weights

Distance-based: fixed geographic extent, variable number of neighbors
Smaller distance captures finer spatial structure, larger distance captures broader structure
Semivariogram
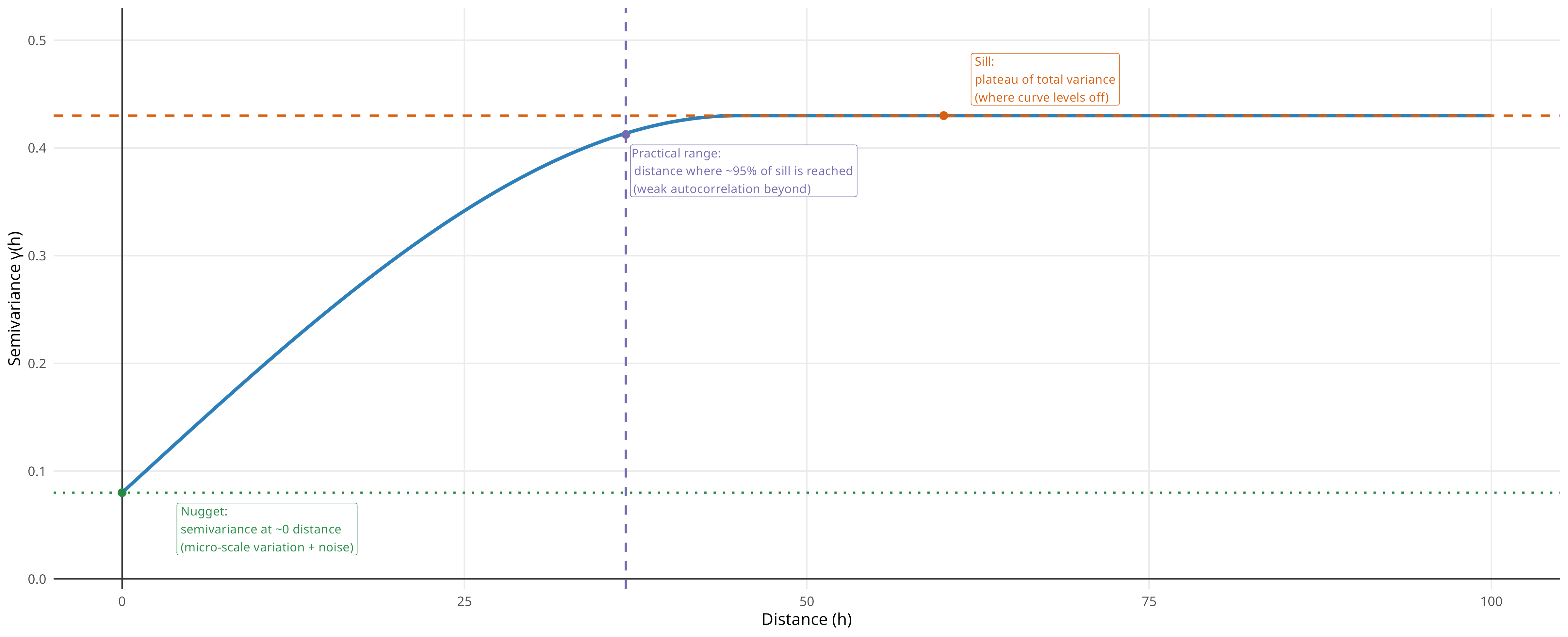
Semivariogram represents dissimilarity of observations as a function of distance, capturing spatial structure across distances
SSVR (Spatially Structured Variance Ratio)
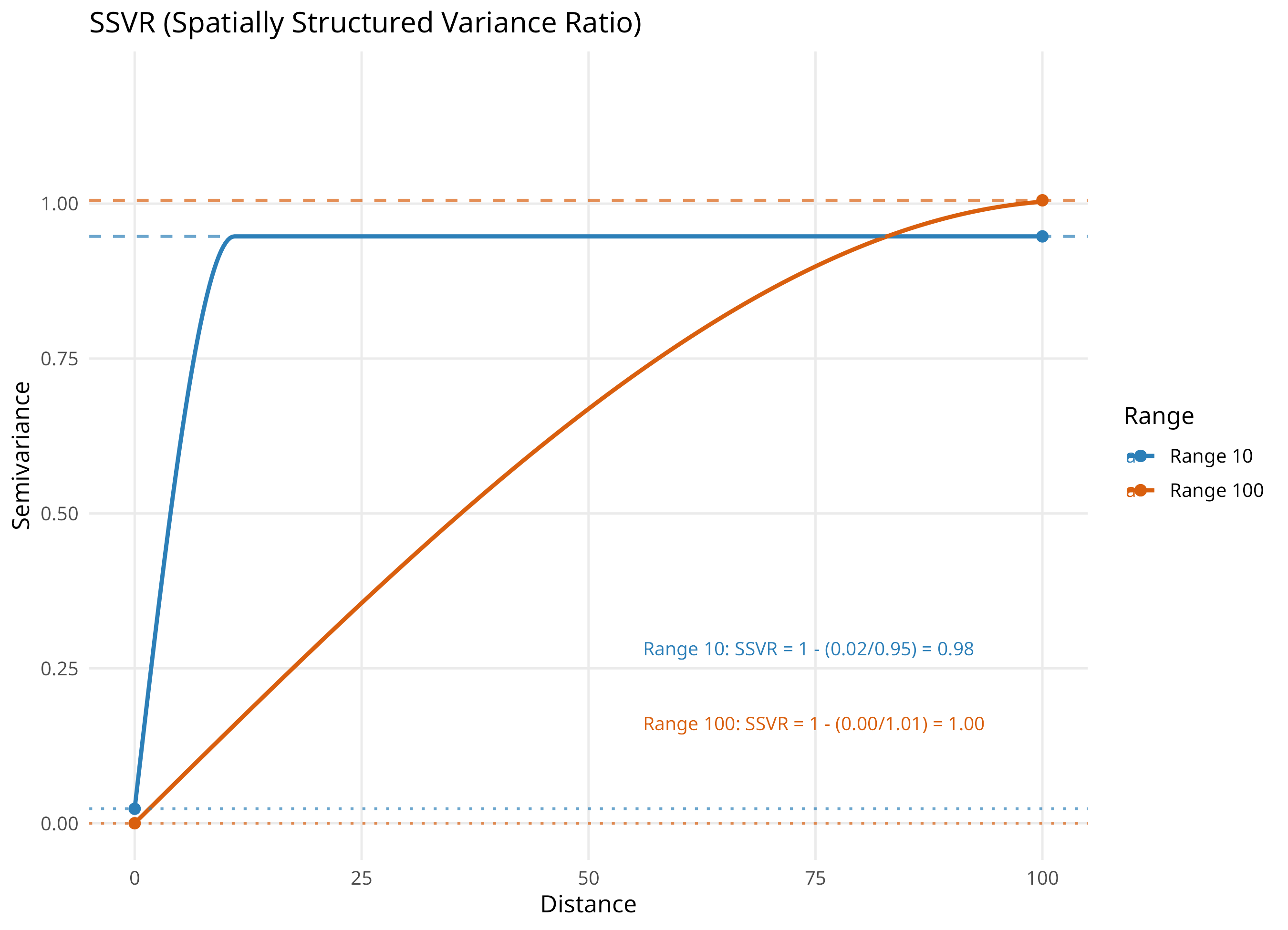
 SSVR (Kerry and Oliver, 2008): share of spatially structured variance
SSVR (Kerry and Oliver, 2008): share of spatially structured variance
An overall summary; ignores where in distance the structure occurs
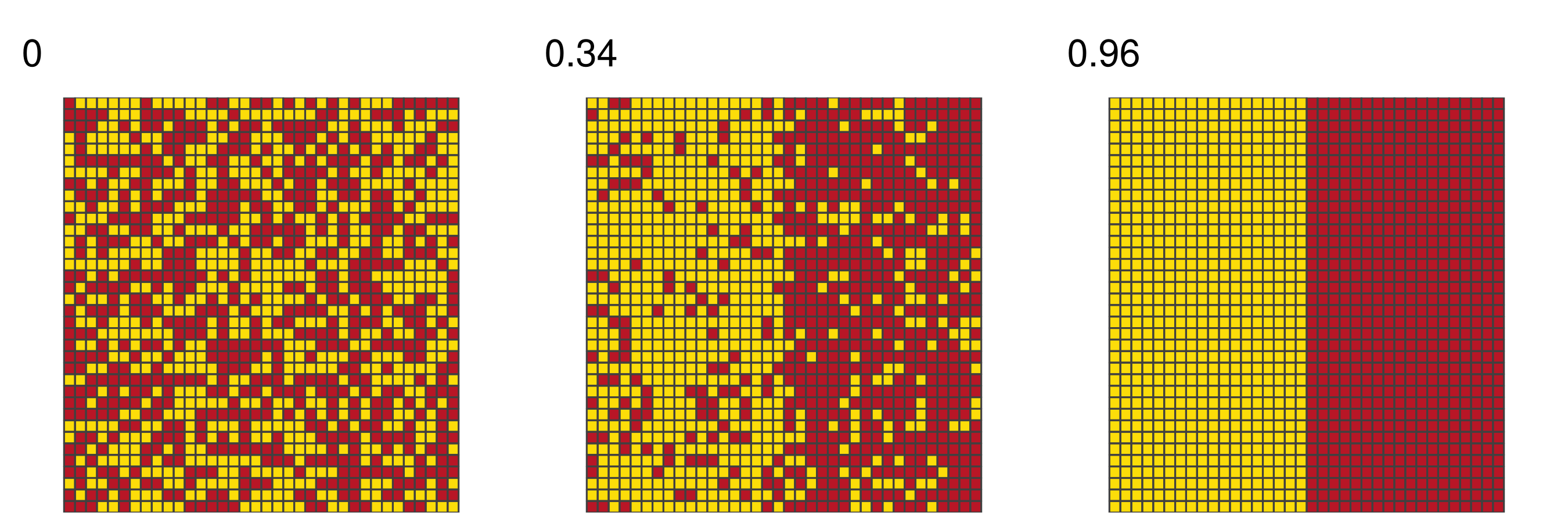
Closer to 1 means stronger spatial structure
AUC (Area Under the Variogram Curve)
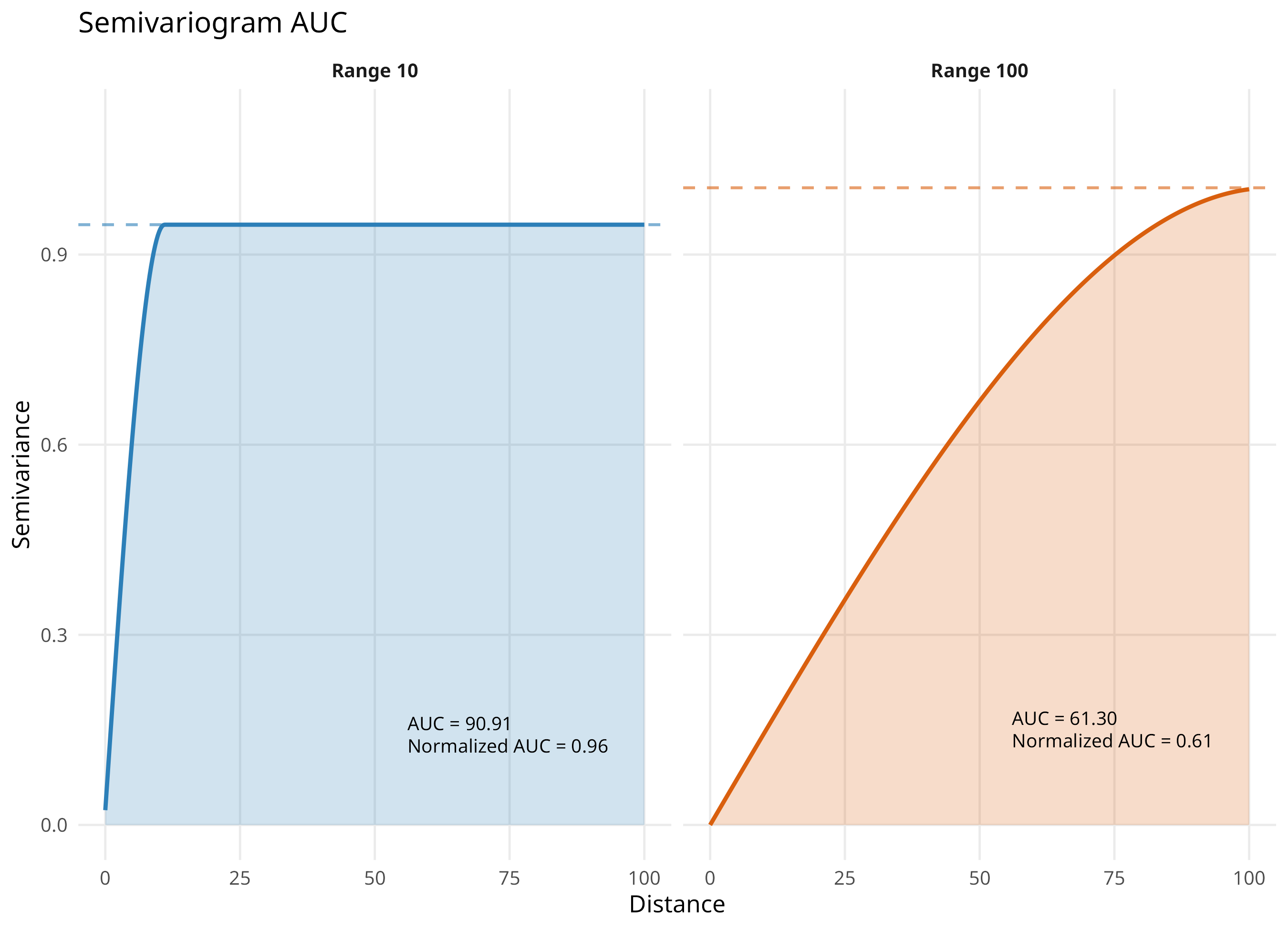
 AUC of variogram (Poggio et al., 2019)
AUC of variogram (Poggio et al., 2019)
Integrates the spatial structure across distances, providing a single summary metric
Larger AUC values indicate stronger spatial dependence
- Integrates the variogram across distances
- Larger AUC values indicate stronger spatial dependence
Setup
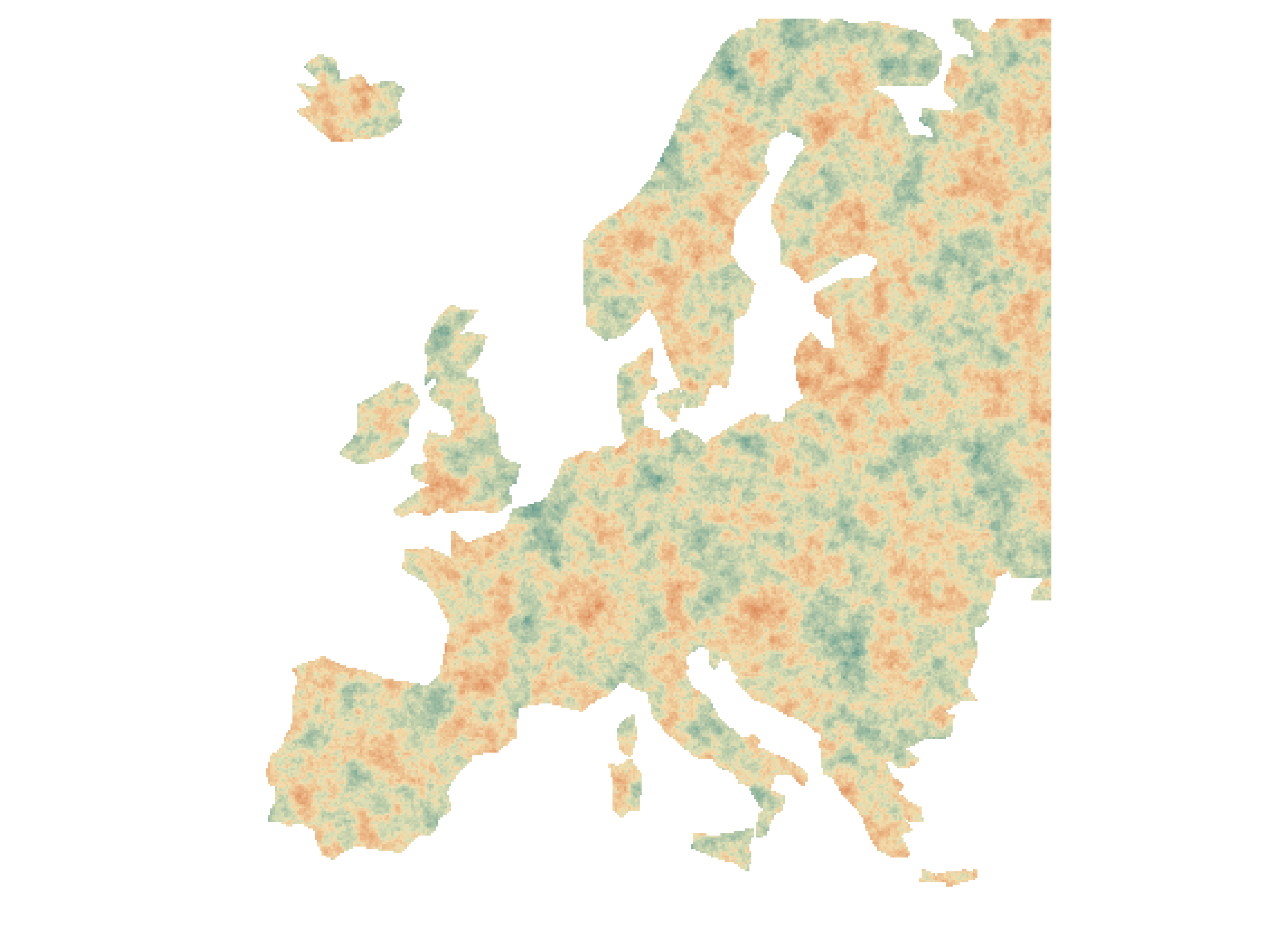
Simulation setup
Simulated rasters with three autocorrelation ranges (10, 50, 100 units)
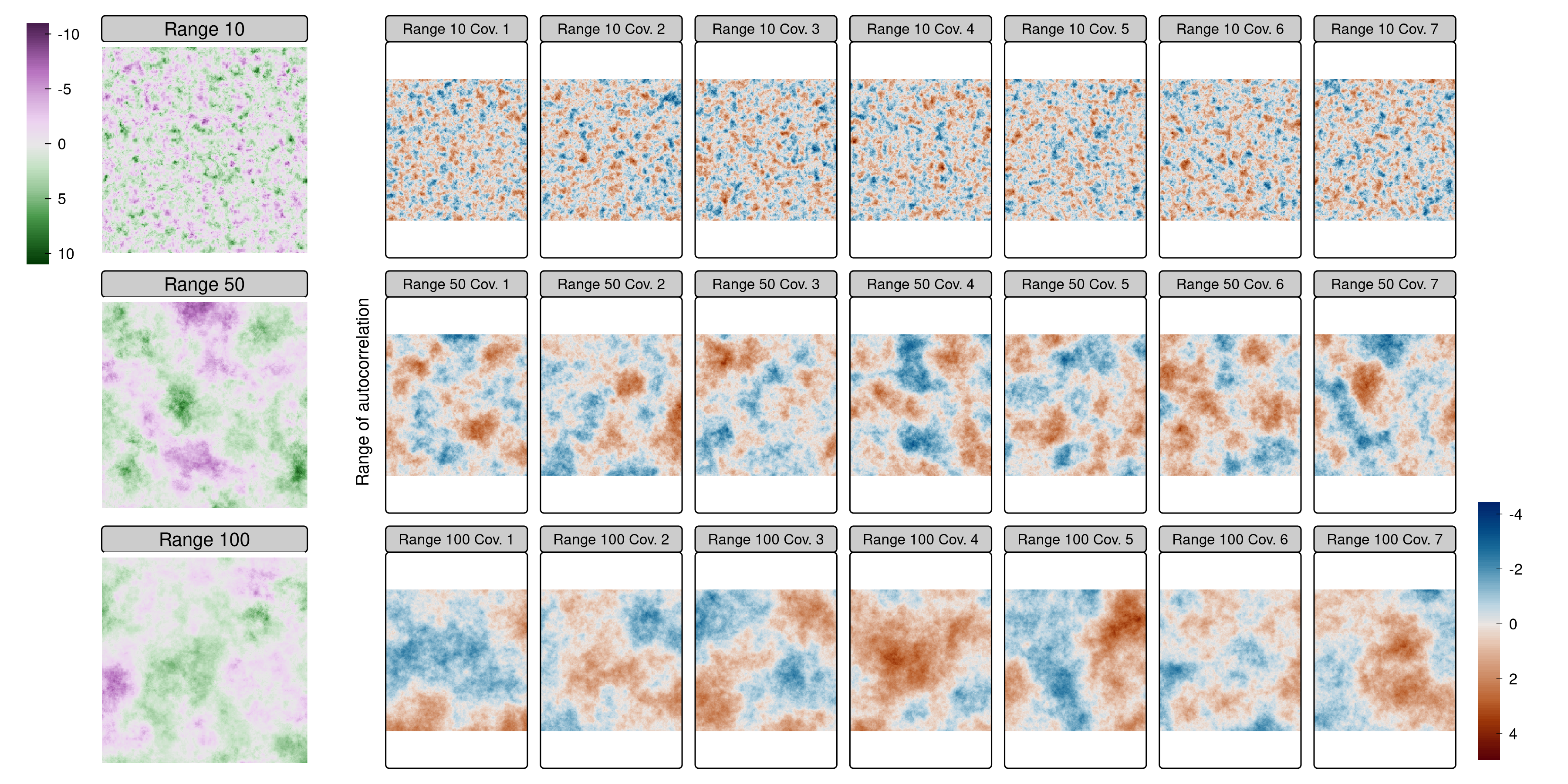
Modeling setup
Random forest models fitted on samples of 500 points
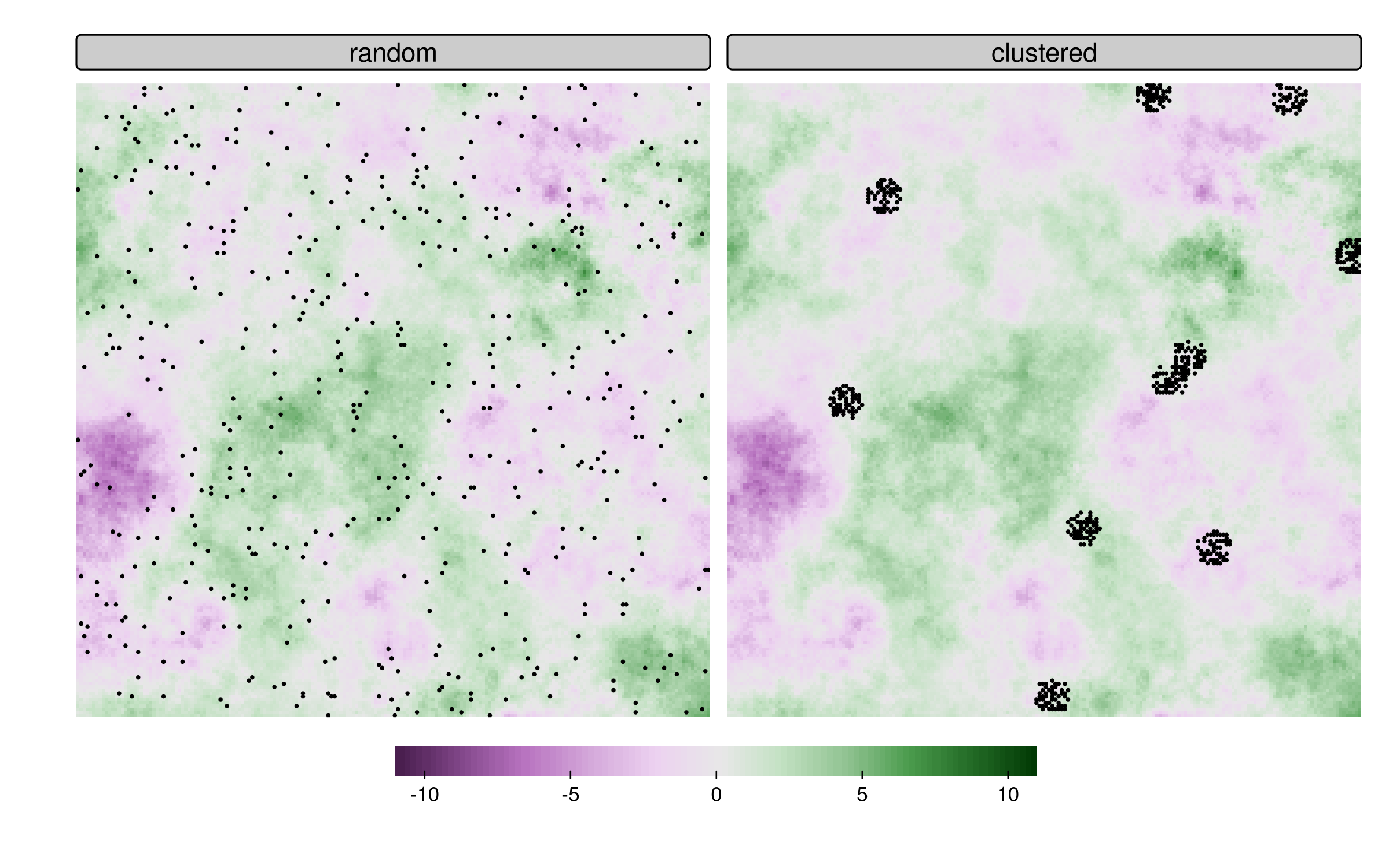
Validation setup
- Complete validation raster
- Four test set sizes
- Two test set sampling types
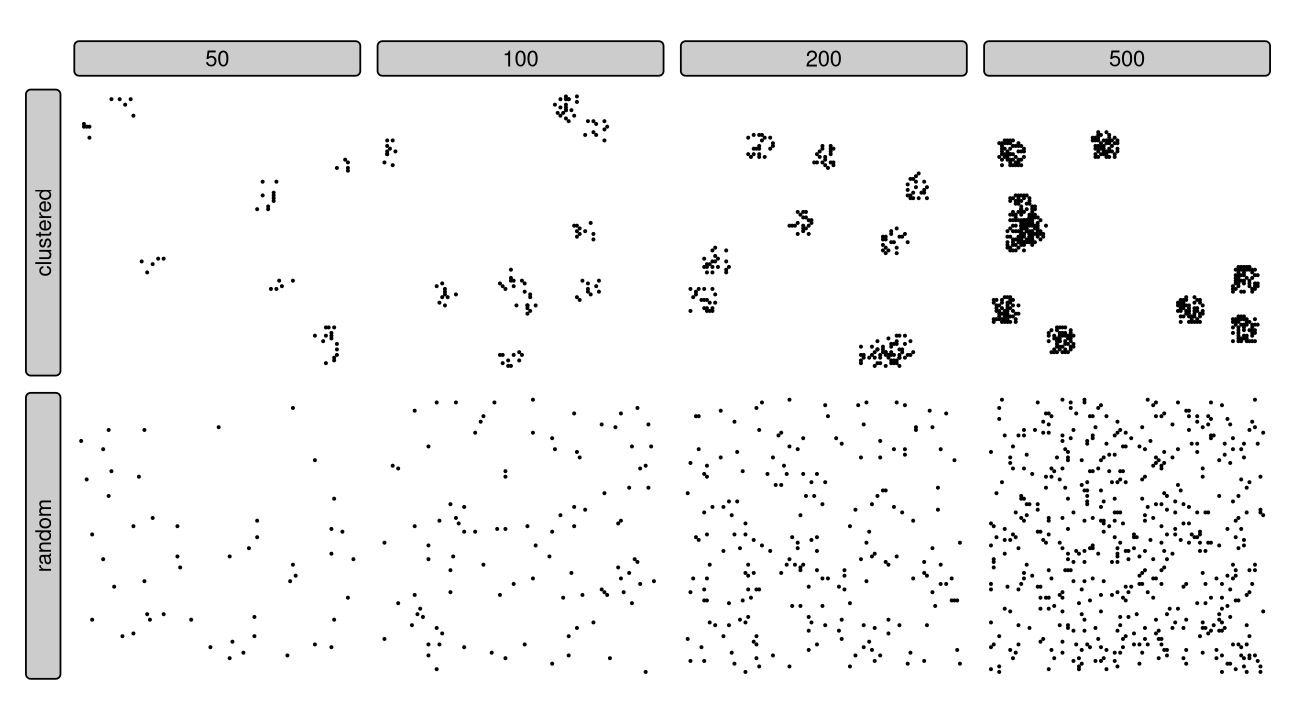
Diagnostic metrics:
- Moran’s I (kNN) (k=5)
- Moran’s I (distance) (from 0 to 100 units)
- SSVR
- AUC
- RMSE
\[ RMSE = \sqrt{\frac{1}{n} \sum_{i=1}^{n} (y_i - \hat{y}_i)^2} \]
Results
Single example showcase
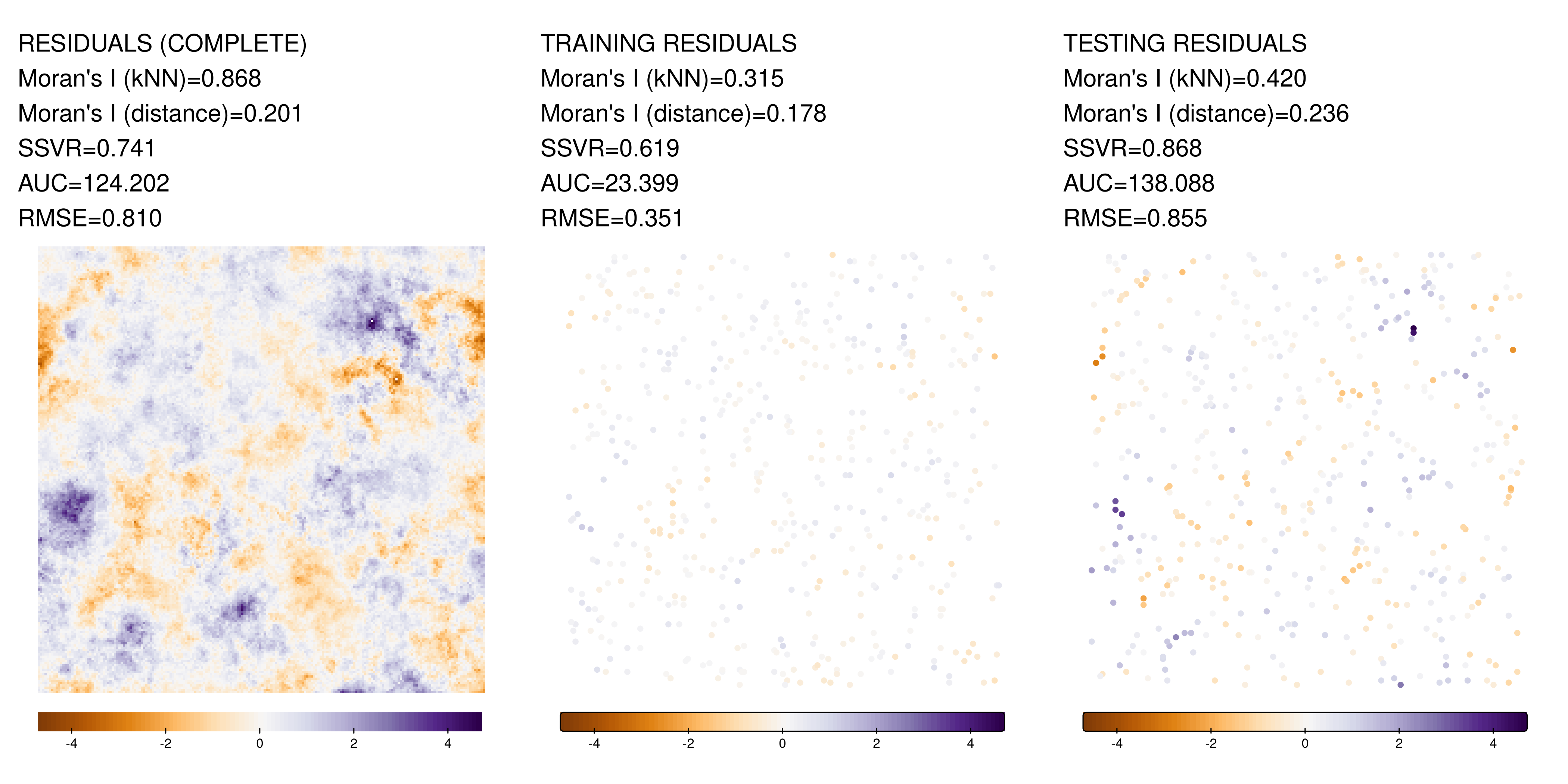
Relationship between metrics for different scopes
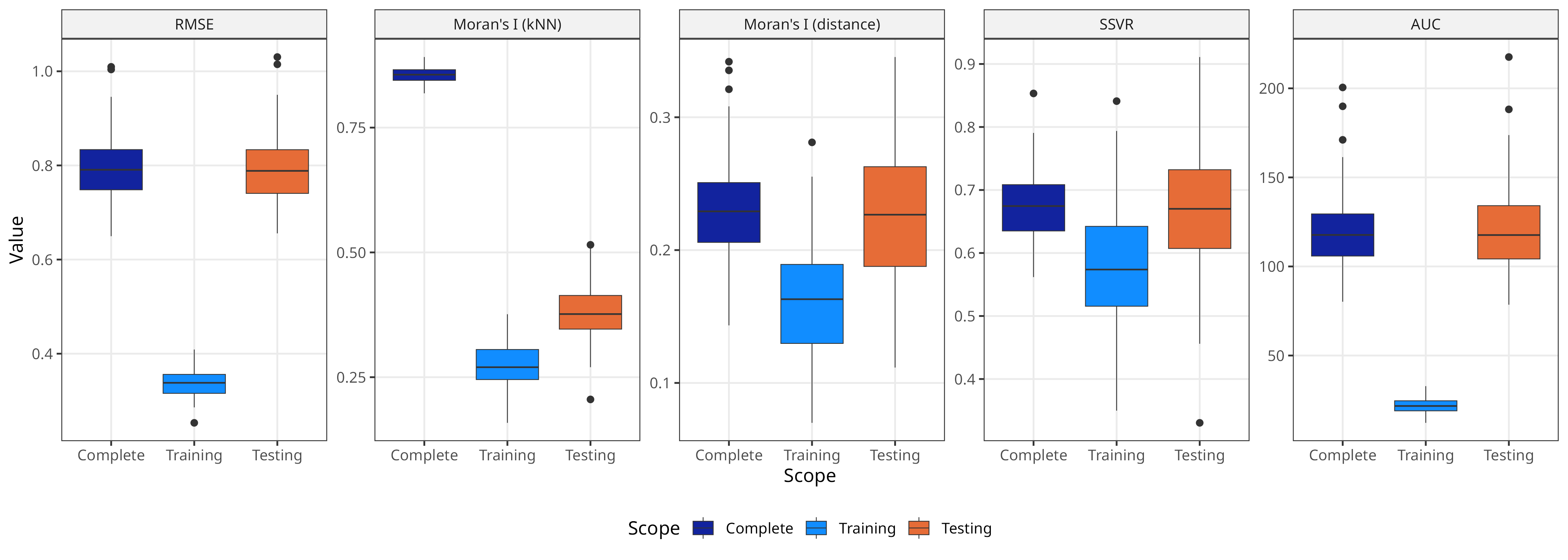
Metrics calculated on testing residuals are mostly comparable to those calculated on complete rasters
An exception is Moran’s I (kNN), with testing values much lower than complete values
Impact of test set size
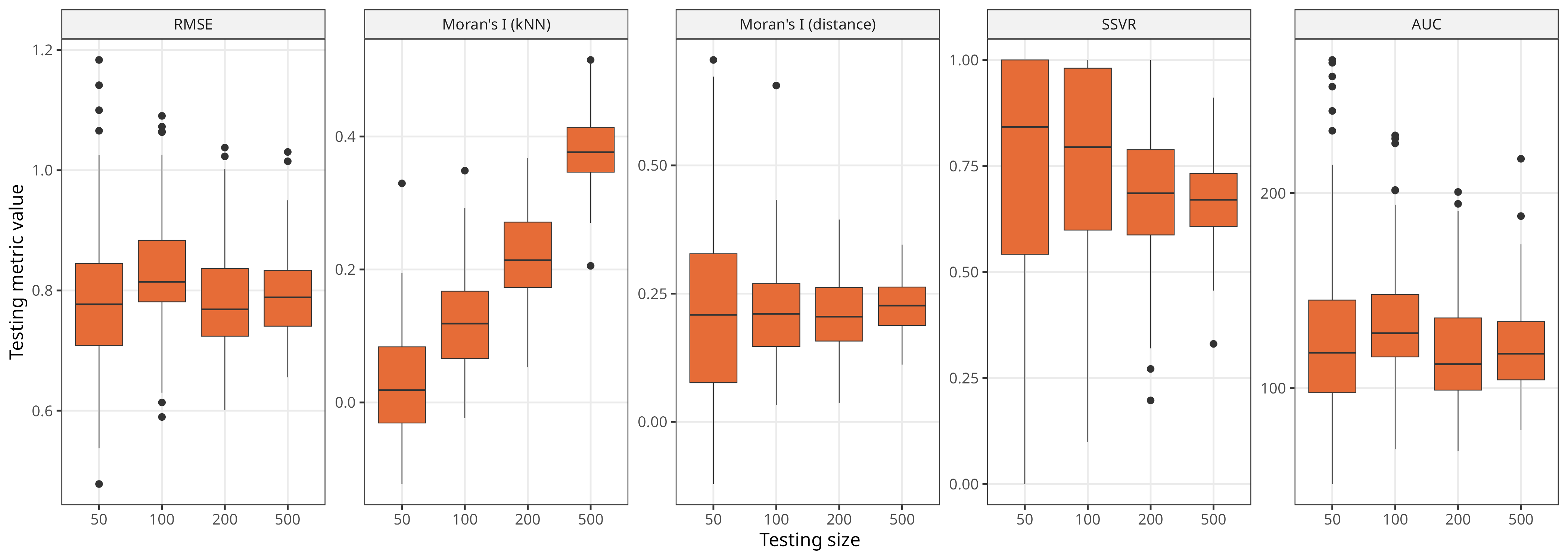
For most metrics, variability decreases as test set size grows, while mean values stay stable
Exception is again Moran’s I (kNN), which shows a strong increase in mean values with larger test set size
Random vs clustered sampling
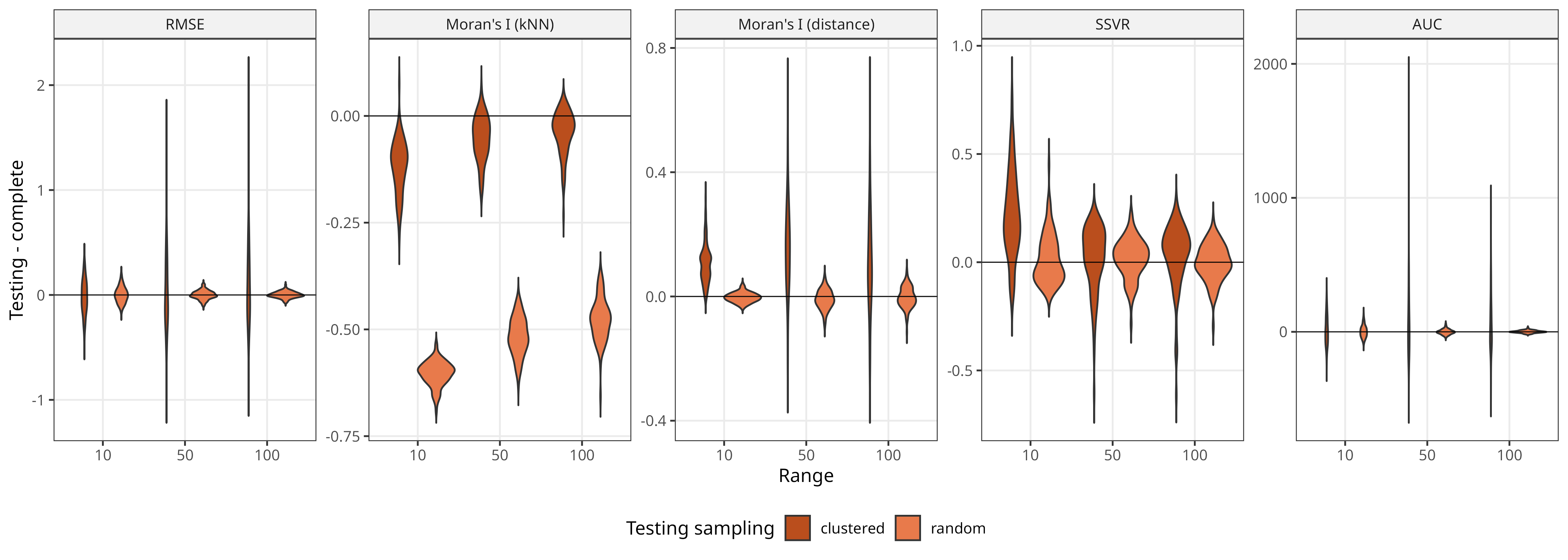
Clustered sampling affects all metrics, leading to higher variability and often incorrect mean values.1
Correlation with RMSE
Based on the results for range = 100 and testing size = 500 with random sampling
| Testing | Complete | |
|---|---|---|
| Moran's I (kNN) | 0.36 | 0.43 |
| Moran's I (distance) | 0.25 | 0.37 |
| SSVR | 0.39 | 0.32 |
| AUC | 0.98 | 0.90 |
Variogram AUC shows the strongest correlation with RMSE – it is a multiscale summary of spatial structure, it captures the overall spatial autocorrelation of residuals. This is is closely related to model performance
Comparison of metrics
Comparison of metrics
| Metric | What it tells | Pitfall |
|---|---|---|
| Moran’s I (kNN) | Autocorrelation at some distance (?) (under fixed neighbor count) | Highly sensitive to k, sample size, and sampling design |
| Moran’s I (distance) | Autocorrelation within a chosen distance range | Sensitive to distance thresholds; more computationally demanding |
| SSVR | Share of spatially structured variance | Requires variogram fit; unstable with small n |
| Variogram AUC | Overall spatial structure across distances | Can track RMSE closely (maybe redundant?) |
Summary
Summary
- Assessing spatial autocorrelation in residuals is key for diagnosing model performance and understanding spatial structure in residuals
- Multiple metrics exist, each with different strengths and limitations
- They measure different aspects of spatial structure
- Report calculation parameters (e.g., k in kNN Moran’s I, distance thresholds) for transparency and reproducibility
Contact
Website: https://jakubnowosad.com
Resources
Slides: https://jakubnowosad.com/egu2026 Software: http://jakubnowosad.com/sacmetrics/

Take-home message


- Moran’s I examines autocorrelation at specific scales defined by the spatial weights
- SSVR summarizes the proportion of variance that is spatially structured
- Variogram AUC integrates spatial structure across all distances, closely tracking RMSE
